Create Synthetic PXMOD Study is an extension of the Create Synthetic KM Study approach. The idea is that synthetic model curves can be generated for many different "tissues", ie. combinations of model parameters, at once. These curves are compiled into images, so that a synthetic study is generated which has a different combination of model parameters in each pixel.
For instance, FDG TACs can be simulated by loading a FDG .km file, selecting the FDG 2- tissue compartment model and using the parameter ranges
Parameter |
From |
To |
K1 |
0.05 |
0.1 |
k2 |
0.06 |
0.11 |
k3 |
0.04 |
0.08 |
k4 |
fixed to 0.0 |
|
To enter these parameter ranges select Create Synthetic PMOD Study from the File. The following dialog appears showing the parameters of the current model.
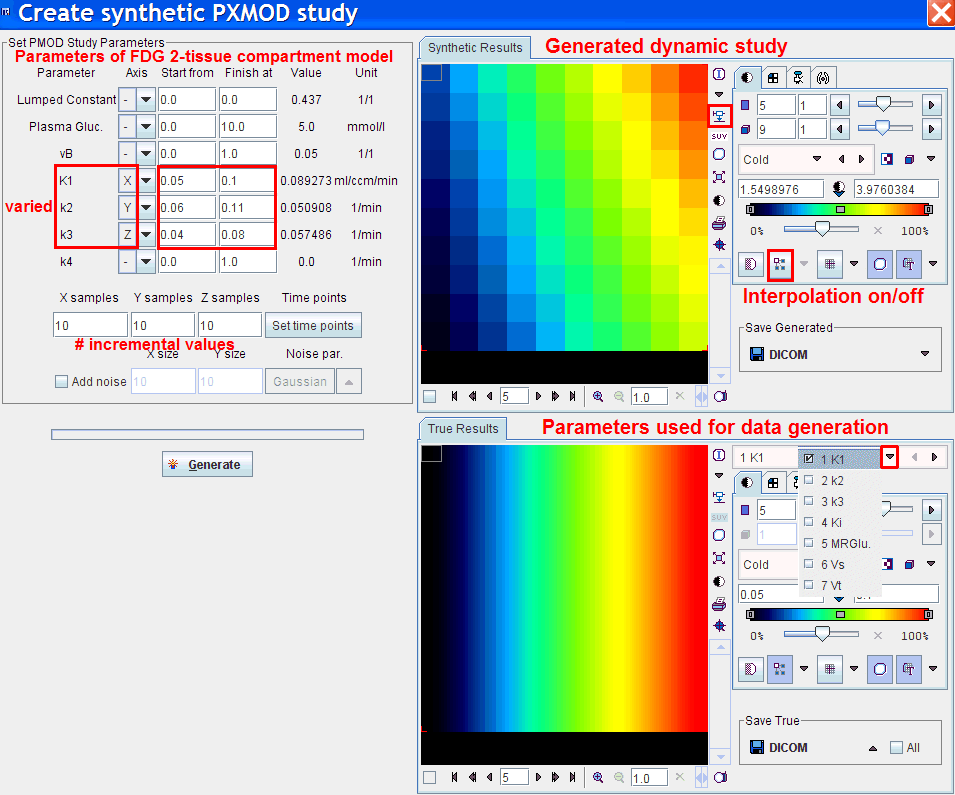
To define the synthetic study the following configurations must be set:
With Generate the model curves of all parameter combinations are generated and the data arranged as a 4-dimensional study. This synthetic study, which is suitable for pixel-wise modeling, is shown in the upper image display. In the example above the interpolation has been disabled to show the 10x10 layout of parameter combinations in plane. Use the Save Generated button to save the study in any supported output format.
The lower image display contains the true parameter images. These generation parameters can be saved using the Save True button. They will be helpful for the analysis of the outcome from a pixel-wise analysis of the synthetic study.