The MP4A RLS (Nagatsuka) model has been developed for the non-invasive quantification method (RLS) of the acetylcholinesterase (AChE) activity in the human brain from measurements with the 11C-MP4A acetylcholine analog. In contrast to reference methods for receptor tracers which use a reference devoid of specific binding, the present method uses a reference with very high AChE activity.
By applying the method of Blomqvist, the following multi-linear equation is derived
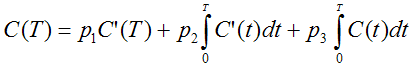
C(t) is the TAC from a cortical target region, and C'(t) the TAC from the reference region (striatum or cerebellum). It can be solved using multi-linear regression, yielding three regression coefficients from which three parameters of interest can be calculated:
R1 = K1/K'1 = p1, the delivery in the target region relative to the reference;
k2 = -p3 -p2/p1 , the rate of back-diffusion from brain to blood;
k3 = p2/p1 , the rate of tracer hydrolysis by AChE.
Implementation Notes
After switching to the MP4A RLS (Nagatsuka) in PKIN a suitable reference region must be selected. The findings in different publications indicate that cerebellum yields more stable results than striatum, most likely due to the higher impact of motion on the signal from the small striatum than the large cerebellum.
While the regression coefficients represent the fitting parameters, the rate constants are shown in the derived parameter section.
Abstract [48]:
N -[(11)C]methylpiperidin-4-yl acetate ([(11)C]MP4A) is an acetylcholine analog. It has been used successfully for the quantitative measurement of acetylcholinesterase (AChE) activity in the human brain with positron emission tomography (PET). [(11)C]MP4A is specifically hydrolyzed by AChE in the brain to a hydrophilic metabolite, which is irreversibly trapped locally in the brain. The authors propose a new method of kinetic analysis of brain AChE activity by PET without arterial blood sampling, that is, reference tissue-based linear least squares (RLS) analysis. In this method, cerebellum or striatum is used as a reference tissue. These regions, because of their high AChE activity, act as a biologic integrator of plasma input function during PET scanning, when regional metabolic rates of [(11)C]MP4A through AChE (k(3); an AChE index) are calculated by using Blomqvist's linear least squares analysis. Computer simulation studies showed that RLS analysis yielded k(3) with almost the same accuracy as the standard nonlinear least squares (NLS) analysis in brain regions with low (such as neocortex and hippocampus) and moderately high (thalamus) k(3) values. The authors then applied these methods to [(11) C]MP4A PET data in 12 healthy subjects and 26 patients with Alzheimer disease (AD) using the cerebellum as the reference region. There was a highly significant linear correlation in regional k(3) estimates between RLS and NLS analyses (456 cerebral regions, [RLS k(3) ] = 0.98 x [NLS k(3) ], r = 0.92, P < 0.001). Significant reductions were observed in k(3) estimates of frontal, temporal, parietal, occipital, and sensorimotor cerebral neocortices (P < 0.001, single-tailed t-test), and hippocampus (P = 0.012) in patients with AD as compared with controls when using RLS analysis. Mean reductions (19.6%) in these 6 regions by RLS were almost the same as those by NLS analysis (20.5%). The sensitivity of RLS analysis for detecting cortical regions with abnormally low k 3 in the 26 patients with AD (138 of 312 regions, 44%) was somewhat less than NLS analysis (52%), but was greater than shape analysis (33%), another method of [(11)C]MP4A kinetic analysis without blood sampling. The authors conclude that RLS analysis is practical and useful for routine analysis of clinical [(11)C]MP4A studies.