Determination of k2' for Models Assuming a Pre-determined k2' Value
Some of the reference methods require an a priori average value of k2', while other methods such as the MRTM or the SRTM reference methods estimate k2' together with the other parameters.
Averaging k2' from Separate Regional Fits
In response to a request from a PMOD user Dr. Ichise recommended the following approach for calculating k2':
- k2' is the tissue clearance rate from the reference region, e.g., cerebellum. This reference tissue is always a region (ROI), not a voxel for both ROI based PKIN or PXMOD parametric imaging. If you define a reference region, there should be only one correct k2' value for that particular subject (scan).
- Logan in her original formulation of her reference tissue model suggested to determine k2' by using arterial data for a group of subjects and use this mean k2'.
- However, we showed that this k2' can be estimated for each individual without arterial data using MRTM or SRTM (three parameter estimation, one of the three parameters is k2').
- I prefer to use k2' estimated this way for each subject for the subsequent MRTM2 or SRTM2. This would be more accurate than the mean k2' estimated for a group of subjects as above.
- Now the accuracy of k2' estimation depends on the following (see [52]): A) noise in the PET data, B) the magnitude of k2' and C) the ratio of k2'/k2 or k2/k2' determined by MRTM or SRTM.
- The bias and variability of k2' estimation by MRTM is less as k2' is larger and k2'/k2 or k2/k2' ratio is further away from unity. This ratio for fallypride using cerebellum and striatum should be greater than 3, I think. In that case, k2' estimation from cerebellum and striatum (use ROIs) should be minimum (see figs in the paper).
- Even using the ROIs, k2' estimation is affected by noise and hence it is good to run MRTM a few times choosing ROIs with high BPNDareas (say right striatum and left striatum) k2'/k2 ratio is further away from 1) and average the k2' values. This averaging is within the subject and totally different from population average.
- Please use the k2' determined as above for estimation of BPND for the cortical regions. The k2' estimated with cerebellum and cortical regions is not accurate because k2'/k2 is closer to unity.
- One advantage of MRTM over SRTM: To use SRTM, both cerebellum and target must be 1 tissue (1T) kinetics (use of the SRTM for 2T will bias BPND). However, MRTM is good for tracers with 2T kinetics such as Fallypride. The only thing here is that you have to give a t* value.
Estimating k2' using a Fit with Regional Coupling
Alternatively, the a-priori knowledge that k2' should always have the same value can be exploited using coupled fitting in PKIN with SRTM2 as follows:
- Select the SRTM2 model for all regions and fit it with k2' enabled.
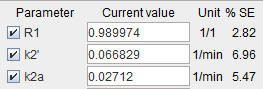
This will result in regionally different k2' values which should be quite similar across the high-binding regions. - Configure a coupled fit with k2' as the common parameter and only include the relevant regions for coupling.
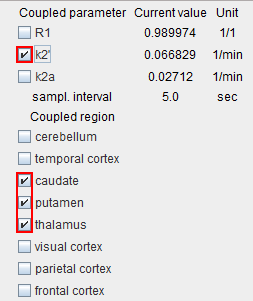
Start the coupled fit which returns a common k2'. This value can be used for pixel-wise fitting or the regional fitting of all TACs - For the regional fitting fix k2' in one of the coupled regions
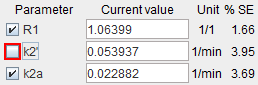
propagate the model to all regions, and then fit all regional TACs. - The same k2' value can also be used for pixel-wise fitting with the PXMOD tool.


