Similar to MA1, Ichise's MA2 analysis method is another alternative technique developed to calculate the total distribution volume of reversible receptor systems with minimal bias. Based on the 2-tissue compartment model equations the following multilinear relationship was derived [34]:
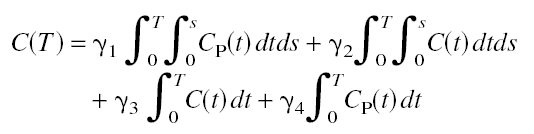
where C(t) represents the tissue time-activity curve, and Cp(t) the plasma activity. A multilinear regression can be performed to calculate the four regression coefficients from the transformed data. The total distribution volume V is then calculated as the ratio ![]() of the first two regression coefficients, and the distribution volume of specific binding by the expression
of the first two regression coefficients, and the distribution volume of specific binding by the expression
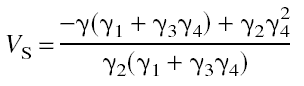
The MA2 method has two advantages:
The authors conclude that for tracers with slow kinetics and low to moderate noise, MA2 may provide the lowest bias while maintaining computational ease.
Implementation Notes
In PKIN, the multilinear regression is performed using a singular value decomposition. Although no equilibration time is required for MA2, there is a Start parameter to disregard early samples from the regression as for the graphical plots and MA1.
Abstract [34]
"In an attempt to improve neuroreceptor distribution volume (V) estimates, the authors evaluated three alternative linear methods to Logan graphical analysis (GA): GA using total least squares (TLS), and two multilinear analyses, MA1 and MA2, based on mathematical rearrangement of GA equation and two- tissue compartments, respectively, using simulated and actual PET data of two receptor tracers, [(18)F]FCWAY and [(11)C]MDL 100,907. For simulations, all three methods decreased the noise-induced GA bias (up to 30%) at the expense of increased variability. The bias reduction was most pronounced for MA1, moderate to large for MA2, and modest to moderate for TLS. In addition, GA, TLS, and MA1, methods that used only a portion of the data (T > t*, chosen by an automatic process), showed a small underestimation for [(11)C]MDL 100,907 with its slow kinetics, due to selection of t* before the true point of linearity. These noniterative methods are computationally simple, allowing efficient pixelwise parameter estimation. For tracers with kinetics that permit t* to be accurately identified within the study duration, MA1 appears to be the best. For tracers with slow kinetics and low to moderate noise, however, MA2 may provide the lowest bias while maintaining computational ease for pixelwise parameter estimation."